
Osteosarcoma is the most common primary malignant bone cancer affecting children and adolescents. Patient survival remains poor due to limited molecular understanding. To address this gap, we integrated single-cell and spatial omics data from >500,000 cells and bulk omics data from >500 patients (manuscript submitted). Built on these rich data resources, we developed OSpaedia.org, a molecular database and analysis platform designed to advance osteosarcoma research through open science.
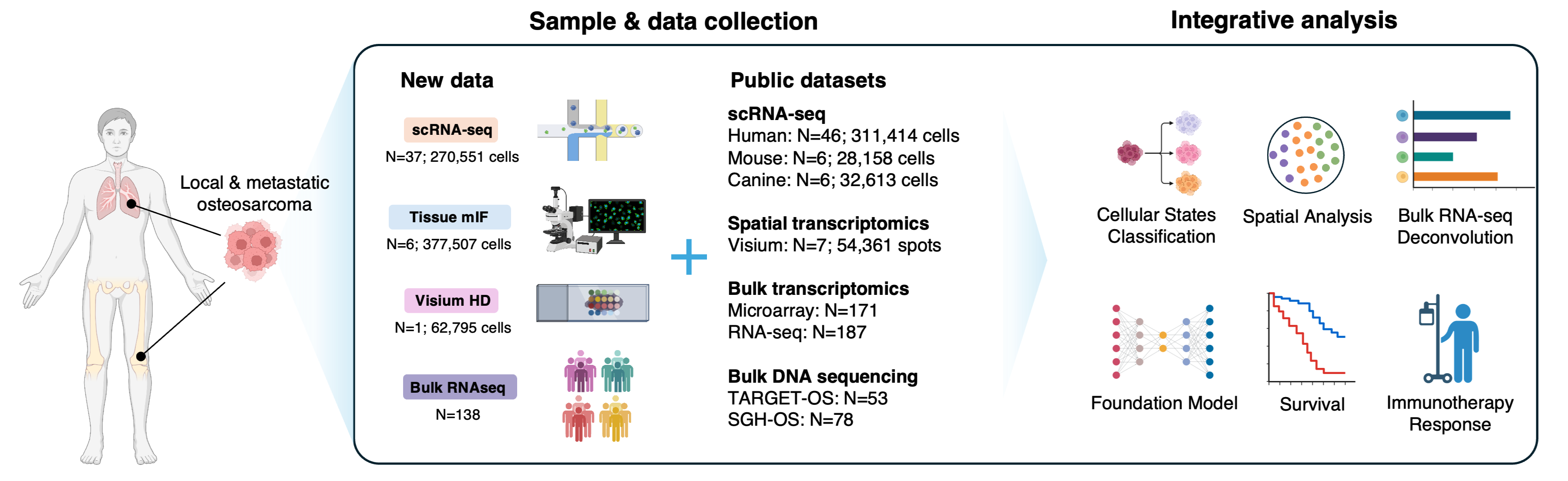
Associated manuscript:
Chen R, Liang H, Ren T, Wang J, Li Y, et al., Integrative single-cell and spatial omics reveal cellular state topography and clinically relevant subtypes in osteosarcoma. 2026, Submitted.
OSpaedia.org is actively growing. Please stay tuned for new datasets and tools.
An integrated single-cell atlas of osteosarcoma for exploring gene expression across cell types and malignant states.
Explore Download Data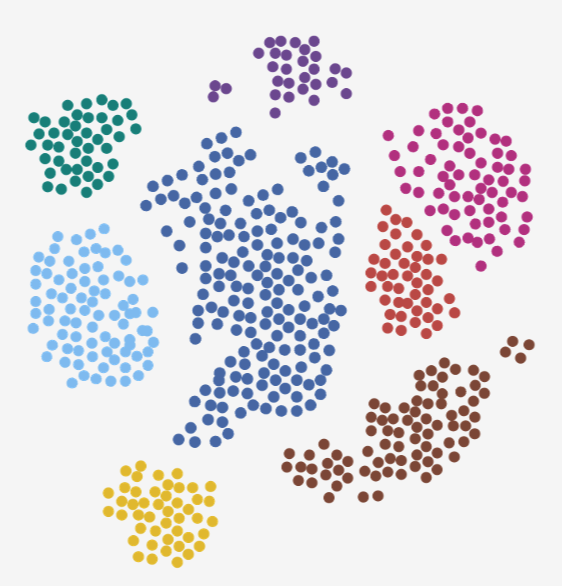
Spatial transcriptomics visualization of tumor, stromal, and immune cell organization.
Explore Download Data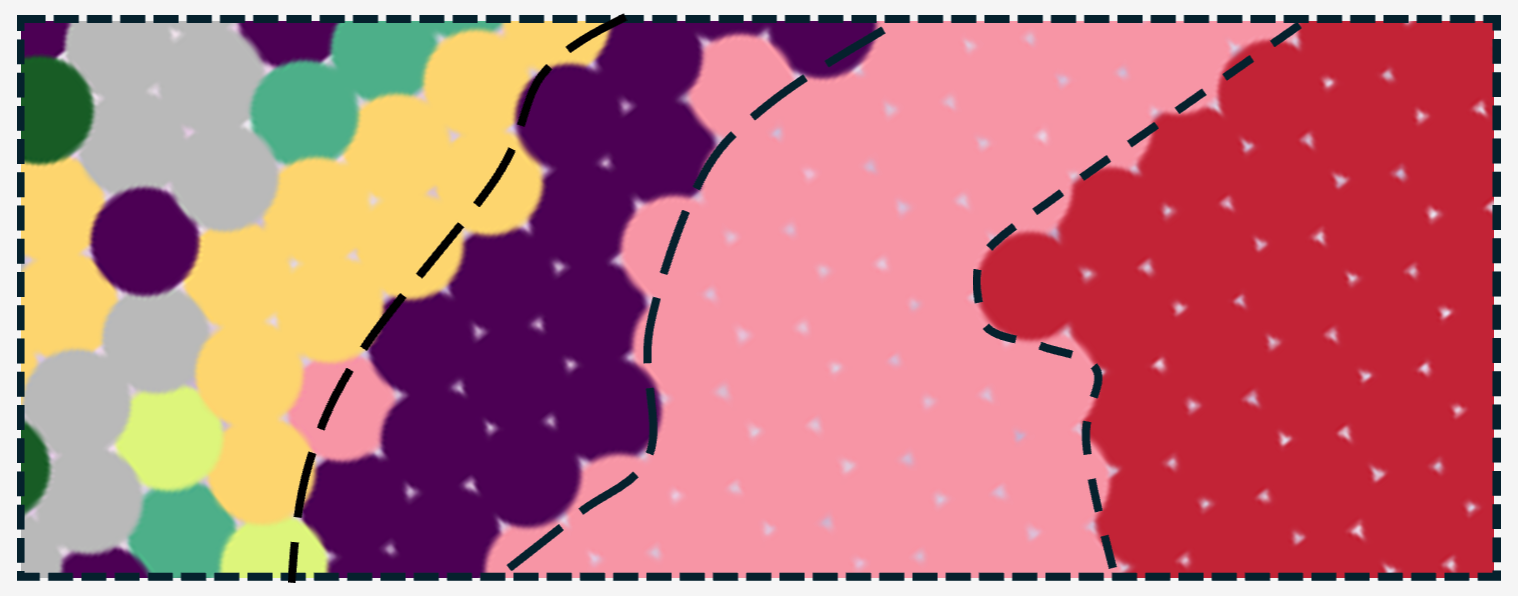
A curated bulk transcriptomic resource with interactive visualization.
Explore Download Data
A bulk RNA-seq–based molecular subtype classifier for osteosarcoma.
Explore on GitHub
A framework for inferring cell-type composition from bulk RNA-seq data.
Explore on GitHub
A fine-tuned single-cell foundation model for automated annotation and batch correction.
Explore on GitHub
We gratefully acknowledge financial support from government funding agencies. We thank the patients and their families for their invaluable contributions. For comments or suggestions, please contact Dr. Quanhua Mu (quanhua.muATpolyu.edu.hk).
Spatial transcriptomic data were retrieved from Code Ocean and Zenodo.